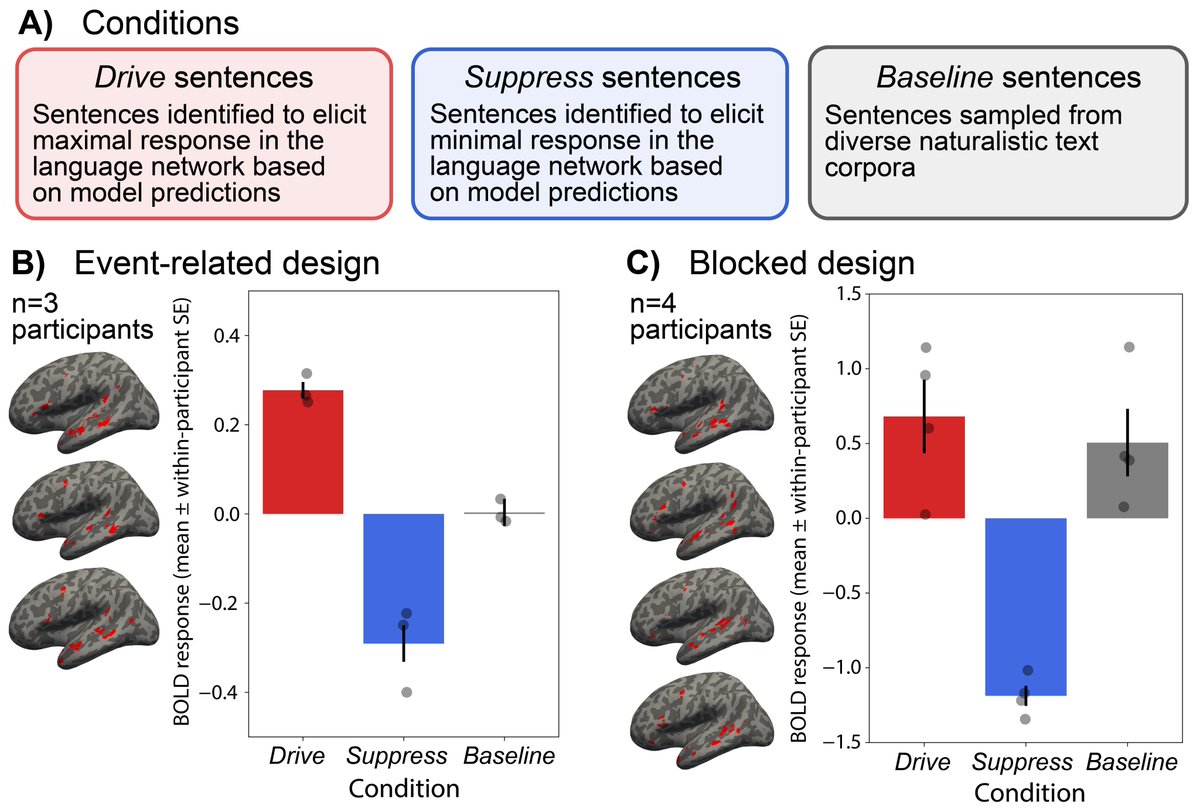
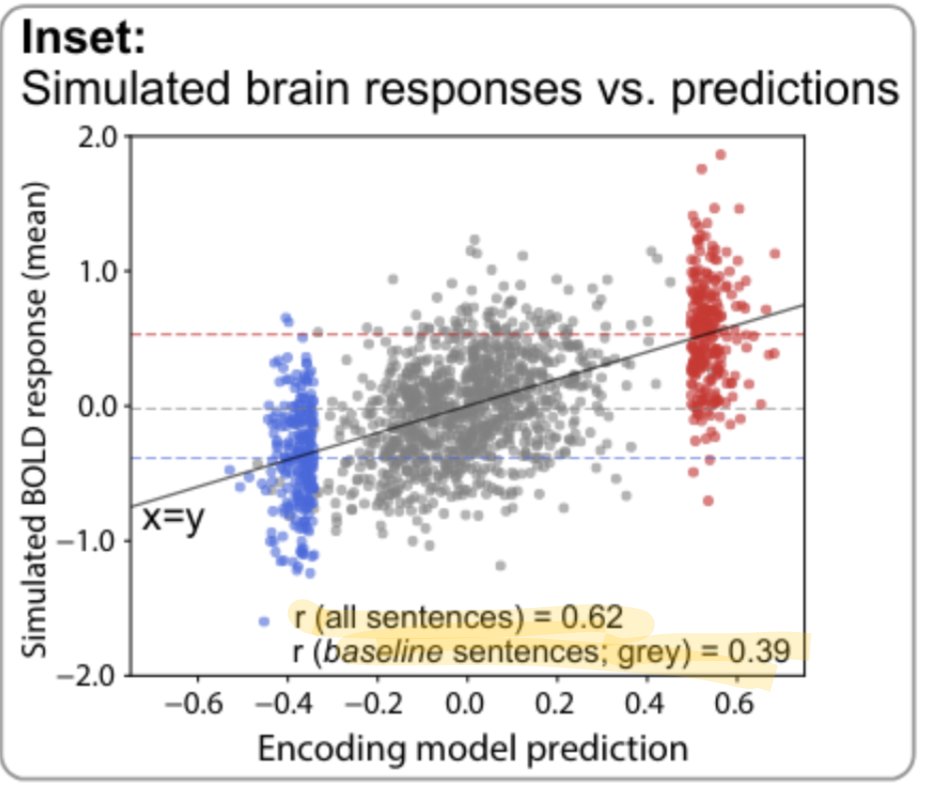
We previously found GPT (2) to be a strong model of the human language system (pnas.org/doi/10.1073/pn).
Author: Martin Schrimpf
Faculty position at EPFL

- Building better (= more behaviorally and neurally aligned) models of vision and language. To build these models, we will use neural recordings from non-human primates and humans as well as human behavioral benchmarks, from the Brain-Score platform [perspective paper, technical paper, CORnet model, VOneNet model, language paper].
- Integrating multimodal representations: I think there is a lot of power in shared invariant representations; creating models that go all the way from pixel input to downstream language representations would allow us to potentially harness shared representations between these two domains, and test if we can better map such a model onto brain hierarchy.
- Towards clinical translation: in my mind, one of the end goals of these brain-modeling efforts is to apply them and improve people’s lives. This could involve helping blind patients (preliminary work), or people with dyslexia.
Wiring Up Vision paper Spotlight at ICLR 2022
Our paper on reducing the number of supervised synaptic updates in computational models of vision was accepted to ICLR as a Spotlight! https://openreview.net/forum?id=g1SzIRLQXMM
The paper improved quite a bit since the preprint I think, we especially made a stronger connection to Machine Learning by showing that our proposed techniques outperform other approaches to drastically reduce the number of parameters. We retain over 40% ImageNet top-1 performance with only ~3% of parameters relative to a fully-trained network.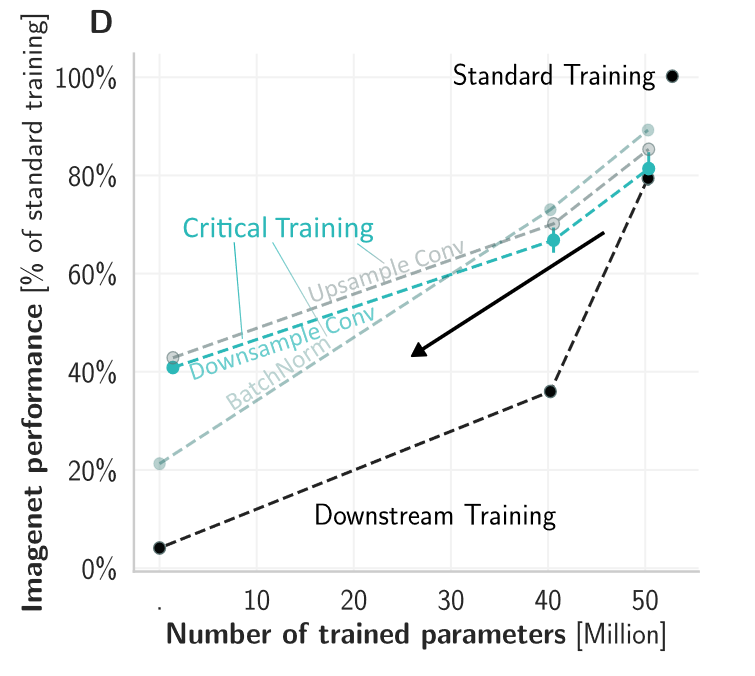
2022 Brain-Score Competition
We are excited to announce that submissions to the 2022 Brain-Score competition are open until February 15, 2022!
The first edition of the Brain-Score Competition proposes to evaluate computational models of primate object recognition in over 30 neuronal and behavioral benchmarks and will award $6,000 to the best submissions over three tracks: overall Brain-Score, V1, and object recognition behavior. In addition, selected participants will be invited to present their work in a Cosyne workshop which will feature some of the leading experts in vision neuroscience and computer vision.
For more information, please visit the competition website, follow Brain-Score on twitter, and join our Slack workspace! Good luck!
ThreeDWorld
Our virtual ThreeDWorld is now public: www.threedworld.org
We provide a fully controllable virtual world striving to be ~photorealistic, based on the Unity engine. ThreeDWorld provides visual and audio rendering with physically realistic behavior that users can interact with through an extensive python API. Check out the code here: github.com/threedworld-mit/tdw
MIT News also wrote a great article summarizing the platform: news.mit.edu
Connecting artificial and biological language processing published at PNAS
Our work modeling the human language system with neural network language models is published in PNAS! https://www.pnas.org/content/118/45/e2105646118
The article received widespread press coverage, e.g. by MIT News, Axios, and Scientific American (Press).
McGovern fellowship and open science prize
I was awarded a Friends of the McGovern fellowship,
and won an Open Science Prize by the Neuro – Irv and Helga Cooper Foundation for my work on Brain-Score.
Teaching Award
I was awarded the Walle Nauta Award for Continuing Dedication in Teaching for the Systems 2 class (Neural Mechanisms of Cognitive Computations) that Mike Halassa and I have been teaching for the past 3 years.
Artificial Neural Networks Accurately Predict Language Processing in the Brain
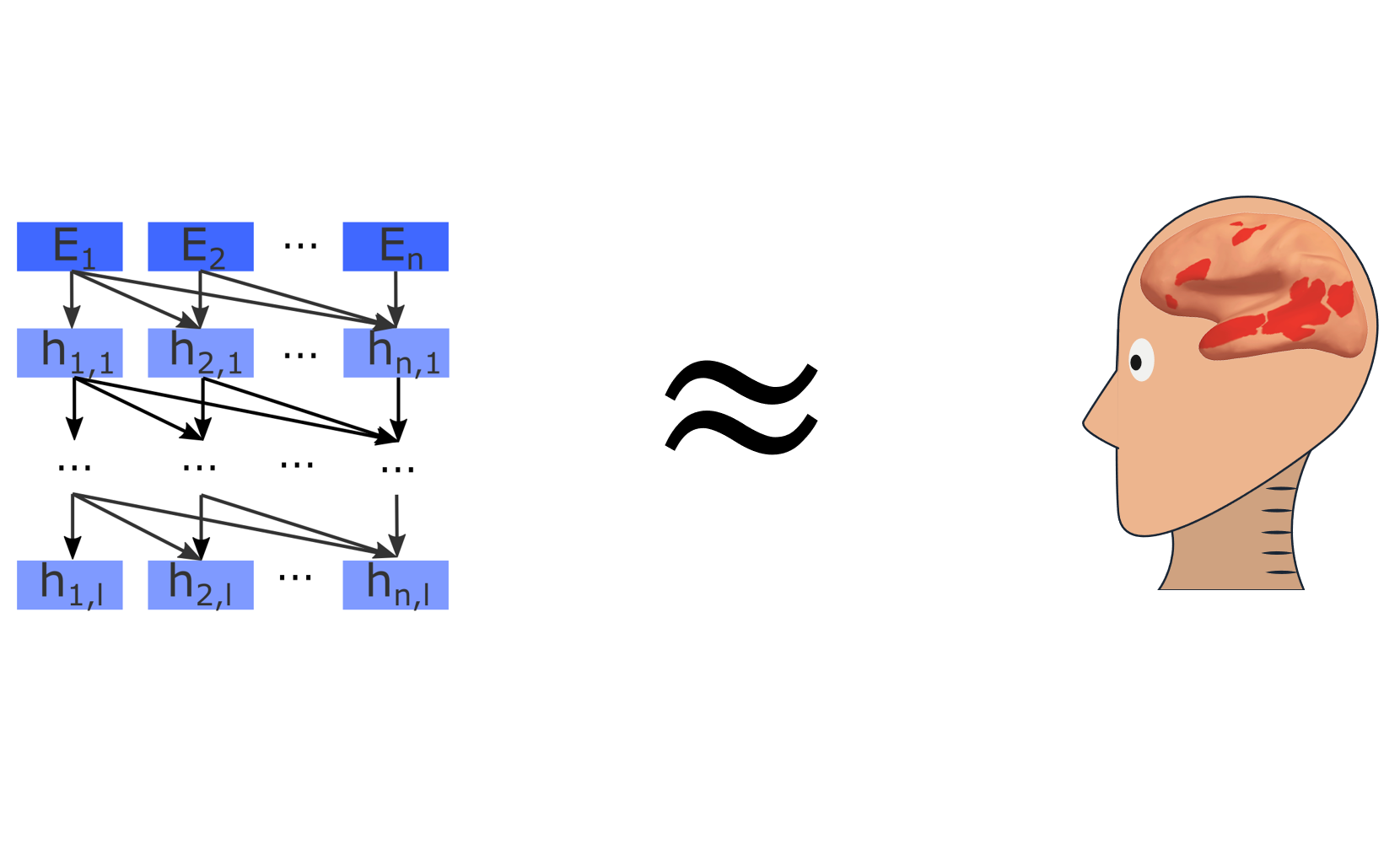 Computational neuroscience has lately had great success at modeling perception with ANNs – but it has been unclear if this approach translates to higher cognitive systems. We made some exciting progress in modeling human language processing https://www.biorxiv.org/content/10.1101/2020.06.26.174482v1.
Computational neuroscience has lately had great success at modeling perception with ANNs – but it has been unclear if this approach translates to higher cognitive systems. We made some exciting progress in modeling human language processing https://www.biorxiv.org/content/10.1101/2020.06.26.174482v1.
This work is the result of a terrific collaboration with Idan A. Blank, Greta Tuckute, Carina Kauf, Eghbal A. Hosseini, Nancy Kanwisher, Josh Tenenbaum and Ev Fedorenko.
Work by Ev Fedorenko and others has localized the language network as a set of regions that support high-level language processing (e.g. https://www.sciencedirect.com/science/article/pii/S136466131300288X) BUT the actual mechanisms underlying human language processing have remained unknown.
To evaluate model candidates of 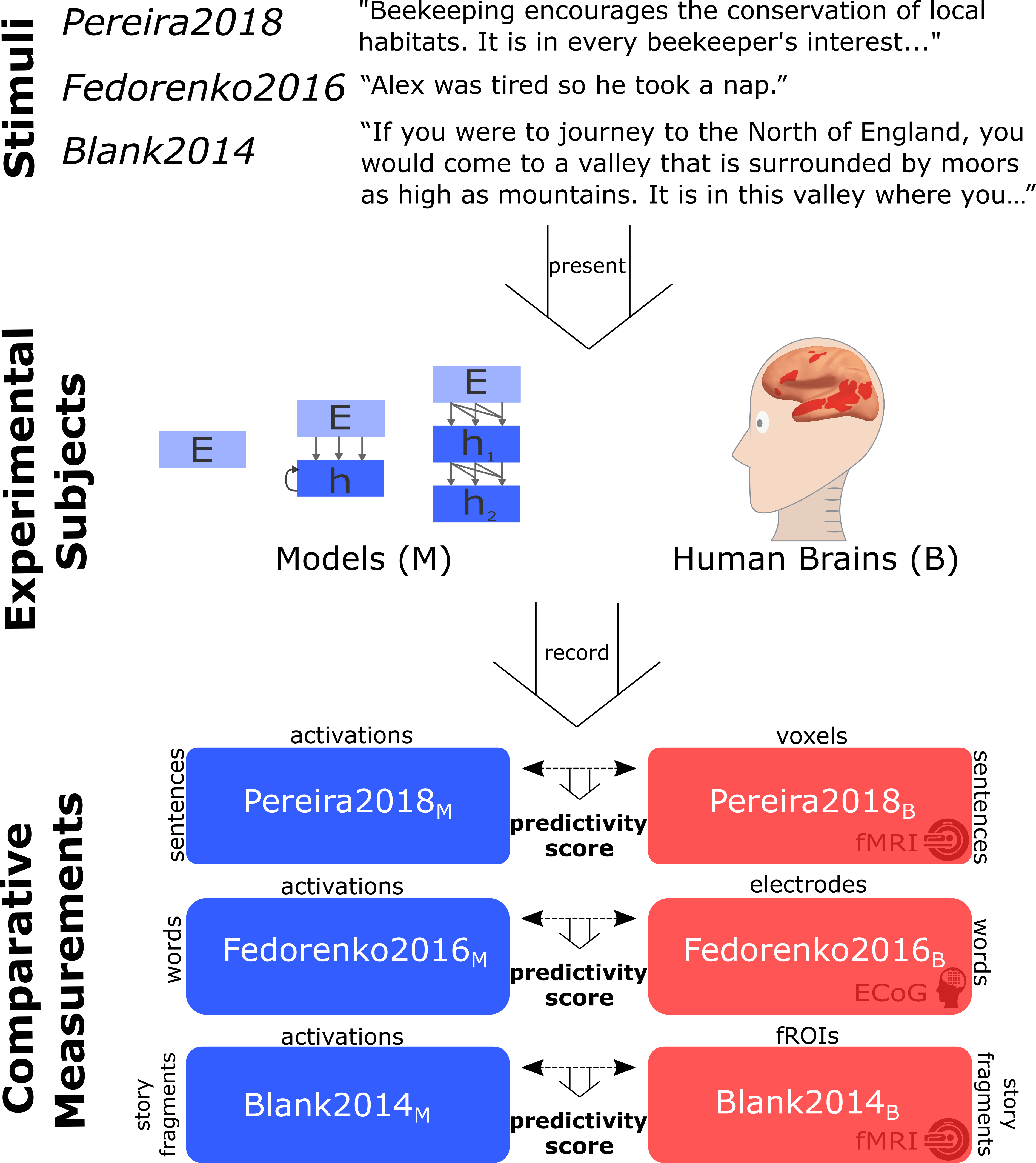 mechanisms, we use previously published human recordings: fMRI activations to short passages (Pereira et al., 2018), ECoG recordings to single words in diverse sentences (Fedorenko et al., 2016), fMRI to story fragments (Blank et al. 2014). More specifically, we present the same stimuli to models that were presented to humans and “record” model activations. We then compute a correlation score of how well the model recordings can predict human recordings with a regression fit on a subset of the stimuli.
mechanisms, we use previously published human recordings: fMRI activations to short passages (Pereira et al., 2018), ECoG recordings to single words in diverse sentences (Fedorenko et al., 2016), fMRI to story fragments (Blank et al. 2014). More specifically, we present the same stimuli to models that were presented to humans and “record” model activations. We then compute a correlation score of how well the model recordings can predict human recordings with a regression fit on a subset of the stimuli.
Since we also want to figure out how close model predictions are to the internal reliability of the data, we extrapolate a ceiling of how well an “infinite number of subjects” could predict individual subjects in the data. Scores are normalized by this estimated ceiling.
So how well do models actually predict our recordings? We tested 43 diverse language models, incl. embedding, recurrent, and transformer models. Specific models (GPT2-xl) predict some of the data near perfectly, and consistently across datasets. Embeddings like GloVe do not.
The scores of models are further predicted by the task performance of models to predict the next word on the WikiText-2 language modeling dataset (evaluated as perplexity, lower is better) – but NOT by task performance on any of the GLUE benchmarks. 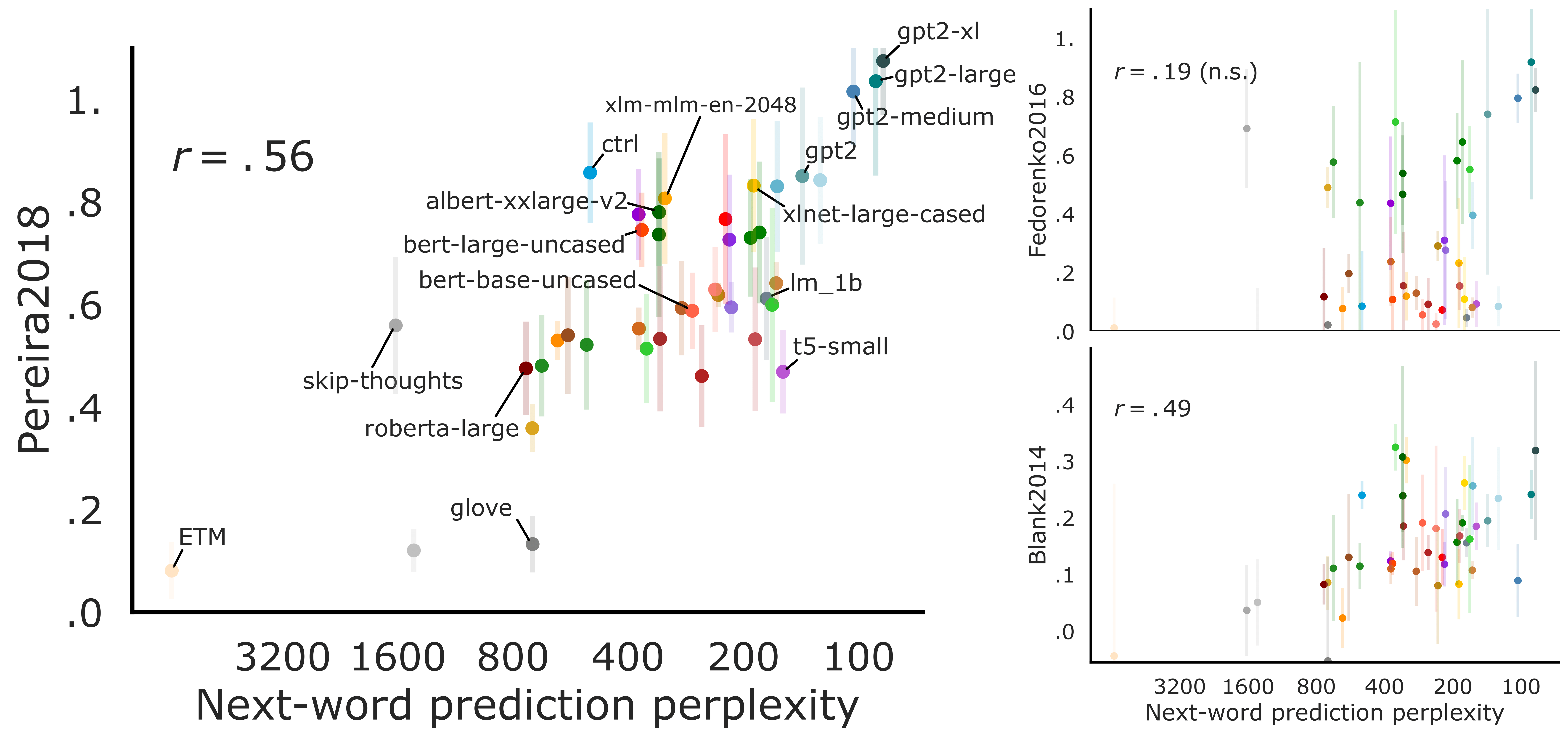
Since we only care about neurons because they support interesting behaviors, we tested how well models predict human reading times: specific models again do well and their success correlates with 1) their neural scores, and 2) their performance on the next-word prediction task.
We also explored the relative contributions to brain predictivity of two different aspects of model design: network architecture and training experience, ~akin to evolutionary and learning-based optimization. (see also this recent work). Intrinsic architectural properties (like size and directionality) in some models already yield representational spaces that – without any training – reliably predict brain activity. These untrained scores predict scores after training. While deep learning is mostly focused on the learning part, architecture alone works surprisingly well even on the next-word prediction task. Critically for the brain datasets, a random embedding with the same number of features as GPT2-xl does not yield reliable predictions.
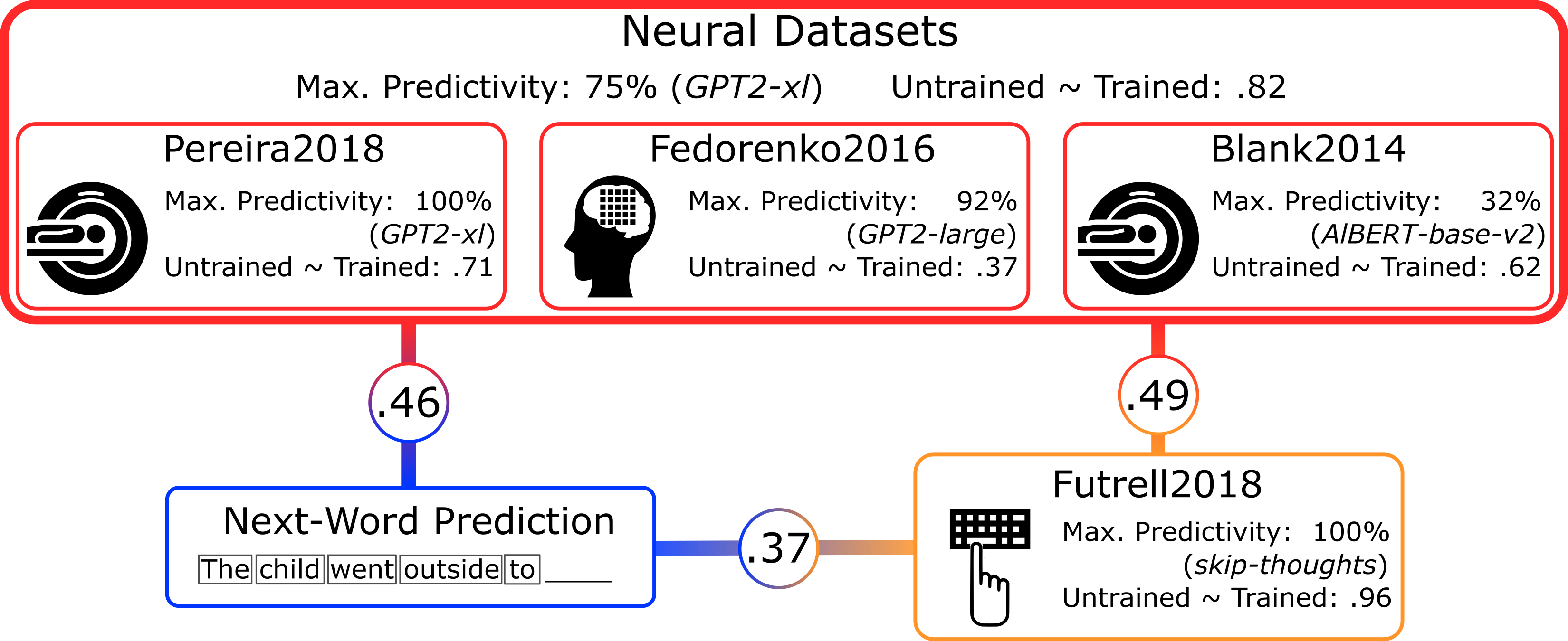 Summary: 1) specific models accurately predict human language data; 2) their neural predictivity is correlated with task performance to predict the next word, 3) and with their ability to predict human reading times; 4) architecture alone already yields reasonable scores. These results suggest that predicting future inputs may shape human language processing, and they enable using ANNs as embodied hypotheses of brain mechanisms. To fuel future generations of neurally plausible models, we will soon release all our code and data.
Summary: 1) specific models accurately predict human language data; 2) their neural predictivity is correlated with task performance to predict the next word, 3) and with their ability to predict human reading times; 4) architecture alone already yields reasonable scores. These results suggest that predicting future inputs may shape human language processing, and they enable using ANNs as embodied hypotheses of brain mechanisms. To fuel future generations of neurally plausible models, we will soon release all our code and data.
Wiring Up Vision: Minimizing Supervised Synaptic Updates Needed to Produce a Primate Ventral Stream
Certain ANNs are surprisingly good models of primate vision, but require millions of supervised synaptic updates — this unbiological development has been the recent focus of many discussions in neuroscience. Is all this training really necessary? We approach this in new work https://www.biorxiv.org/content/10.1101/2020.06.08.140111v1.
Neuroscientists have argued for innate structure with only thin learning on top, i.e. where structure the genome dictates brain connectivity and is leveraged for rapid experience-dependent development. We took first steps at this with more brain-like neural networks.
with only thin learning on top, i.e. where structure the genome dictates brain connectivity and is leveraged for rapid experience-dependent development. We took first steps at this with more brain-like neural networks.
We started from CORnet-S, the current top model on neural and behavioral benchmarks in Brain-Score.org. We first found that variants of this model which are trained for only 2% of supervised updates (epochs x images) already achieve 80% of the trained model’s score.
Even without any updates, the models’ brain predictivities are well above chance. Examining this “at-birth” synaptic connectivity and improving it with a new method “Weight Compression”, we can reach 54% without any training at all
However, to be more brain-lik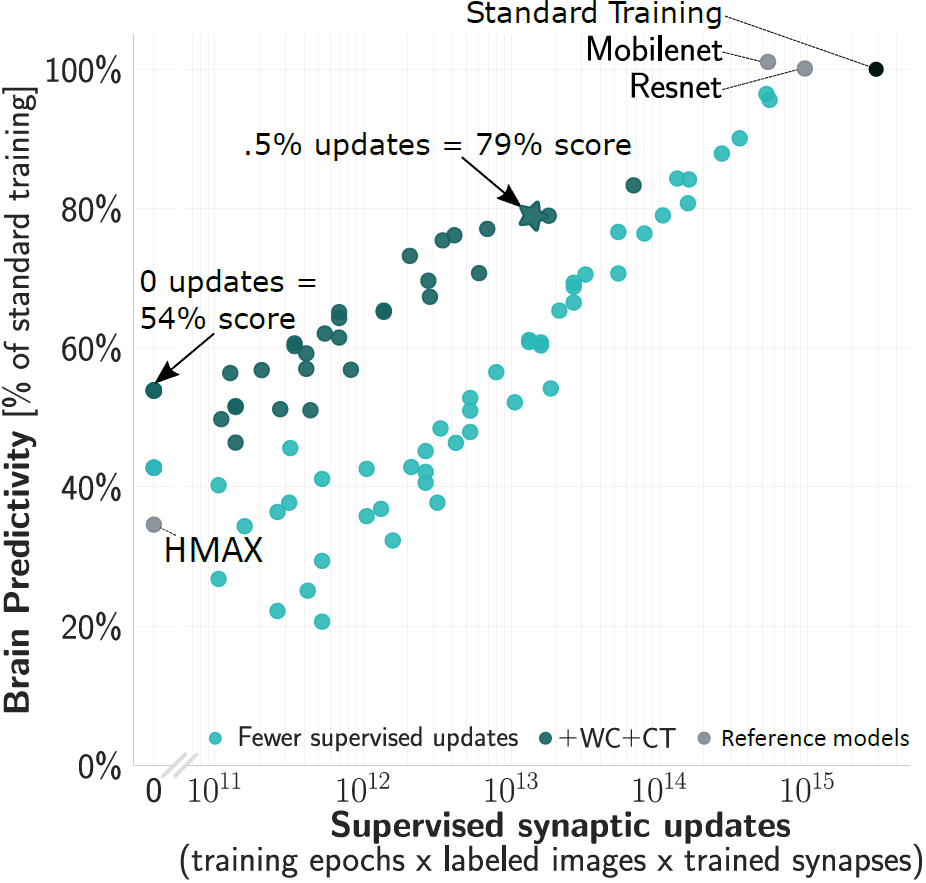 e we require at least some training — but ideally this would not change millions of synapses requiring precise machinery to coordinate the updates. By training only critical down-sampling layers, we achieve 80% when updating only 5% of synapses.
e we require at least some training — but ideally this would not change millions of synapses requiring precise machinery to coordinate the updates. By training only critical down-sampling layers, we achieve 80% when updating only 5% of synapses.
Applying these three strategies in combination (reducing supervised epochs x images + improved at-birth connectivity + reducing synaptic updates), we achieve ~80% of a fully trained model’s brain predictivity with two orders of magnitude fewer supervised synaptic updates.
Taking a step back, we think these are first steps to model not just primate adult visual processing during inference, but also how the system is wired up from an evolutionary birth state encoded in the genome and by developmental update rules. Lots more work to do!
Takeda Fellowship
Very excited to share that I was awarded the Takeda Fellowship on research at the intersection of AI and Health!
My development setup
Since this has been helpful to many colleagues, I thought I would put together my development setup and especially include a small how-to on setting up PyCharm with remote support.
PyCharm remote setup
I have found the remote support of PyCharm to be extremely helpful. It is only available in the professional version, but you get it for free if you’re a student.
The pros are extremely appealing: You can run everything on a remote server while the development experience feels as if it were local. PyCharm takes care of uploading changes, running the remote interpreter (even in debug mode), and displaying figures directly in the IDE.
Follow these steps to setup PyCharm in the same way:
- open File > Settings, navigate to Project > Python interpreter
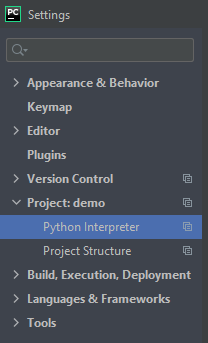
- On the right side of the “Python Interpreter” line, click the gear icon and click “Add…”
- select “SSH Interpreter” and create a new server configuration. Enter the host and your username, click next
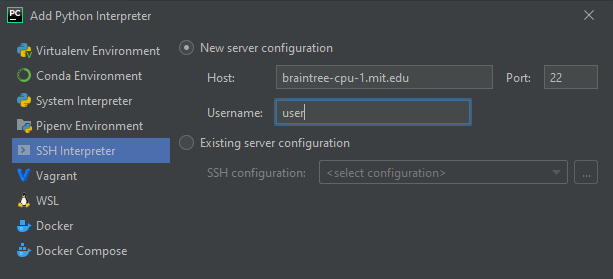
- Enter your password or private key, click Next
- Enter the path to the remote interpreter. I use a local miniconda installation, so for me the path is something like /braintree/home/user/miniconda3/bin/python.
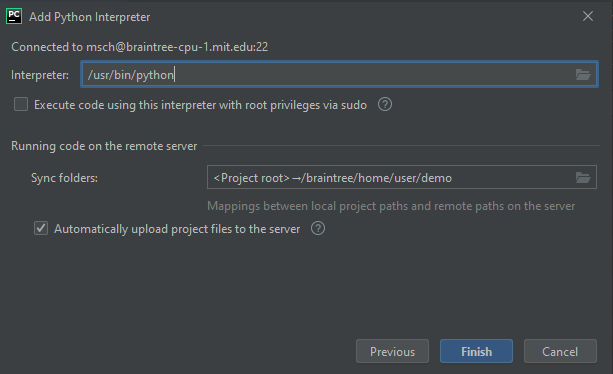
- On the same screen, click the Folder icon to the right of “Sync folders” and input where you want your local files synced. Click Finish
That’s it! You can now program as usual in your IDE and files will automatically be transferred to the remote host. When you run/debug things, they will automatically run on the remote host. (In case it doesn’t sync, make sure your remote path is entered in Settings > Deployment > Mappings.)
Advanced tips
- Running fig.show() with a matplotlib figure will show it directly in the IDE
- For multiple attached projects, you can go to Settings > Build, Execution, Deployment > Deployment > (in view) Mappings to setup the local/remote path pairs
- Make sure to use the integrated Version Control tools. If they don’t show up automatically, check that the directory is recognized under version control in Settings > Version Control
- Use auto-formatting! PyCharm can automatically reformat your code according to a specific (or default) code style
- Use hotkeys! They make it much quicker to accept suggestions, auto-format, run, step into etc. (I use the Eclipse keymap out of habit)
- To install miniconda locally, I usually run wget https://repo.anaconda.com/miniconda/Miniconda3-latest-Linux-x86_64.sh && bash Miniconda3-latest-Linux-x86_64.sh.
Hardware:
- SurfaceBook laptop on a stand
- 2 external 24″ monitors
- Kinesis Freestyle2 split keyboard
- wireless mouse
- external webcam
- wireless headset
- SurfaceDock to bring it all together with minimal daily cable management
Software:
- PyCharm IDE
- KiTTY ssh client
- Keep for todo lists
- OneNote for note-keeping
- PowerPoint for presentations / running plots
- KeePass password manager
- Dropbox for all documents and presentations (except code, but including the password database so that I can access it on my phone through KeePass2Android)
- Github for anything code-related
Oral presentation at NeurIPS
The work led jointly with Jonas Kubilius on “Brain-Like Object Recognition with High-Performing Shallow Recurrent ANNs” was accepted to NeurIPS as an oral! Only 36 (or 0.5%) out of the 6743 submissions were selected as an oral, so we’re very excited to present our ideas how Machine Learning and Neuroscience can interact again in the form of models of the brain.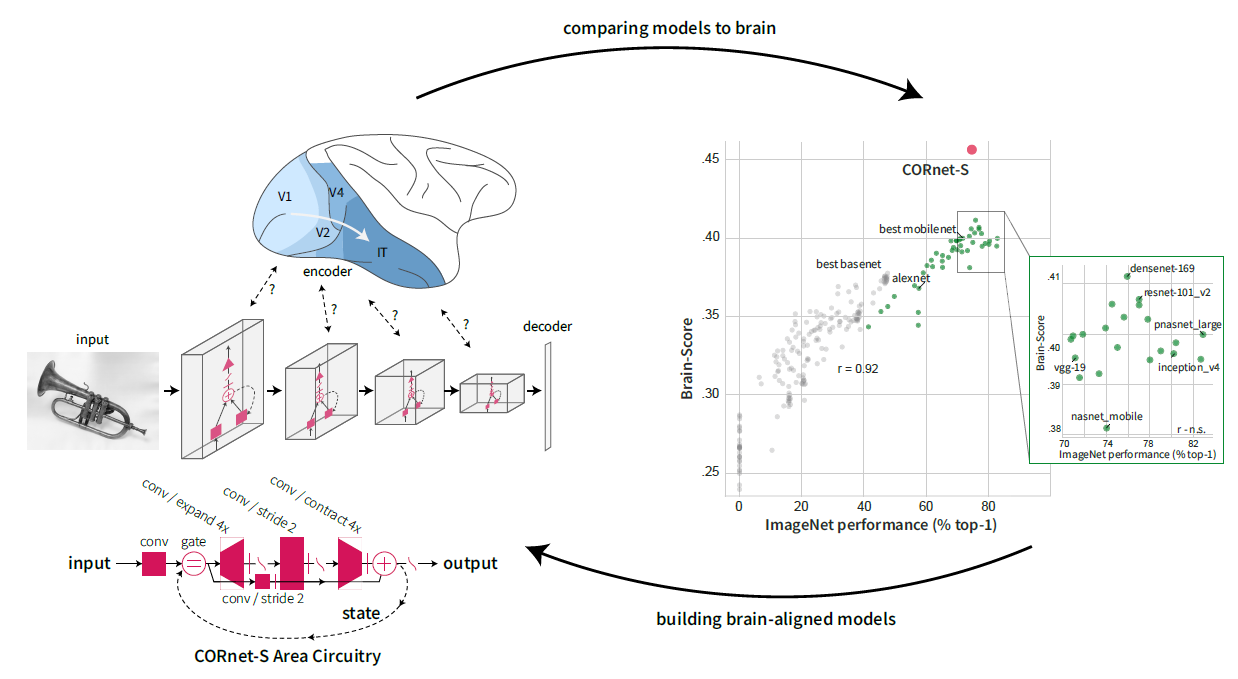
The McGovern institute and Robin.ly have also reported about our findings.
Efficient Neural Stimulation with In-Silico Stimulation, class project
For my class project in Alan Jasanoff’s course Neurotechnology in Action, I experimented with modeling the behavioral effects of microstimulation. I used Afraz et al., Nature 2006 as a reference and started from our recently developed CORnet model (Kubilius*, Schrimpf* et al. NeurIPS 2019). By increasing the activity at parts in the model, I was able to reproduce the behavioral effects reported by Afraz et al. to a first extent. 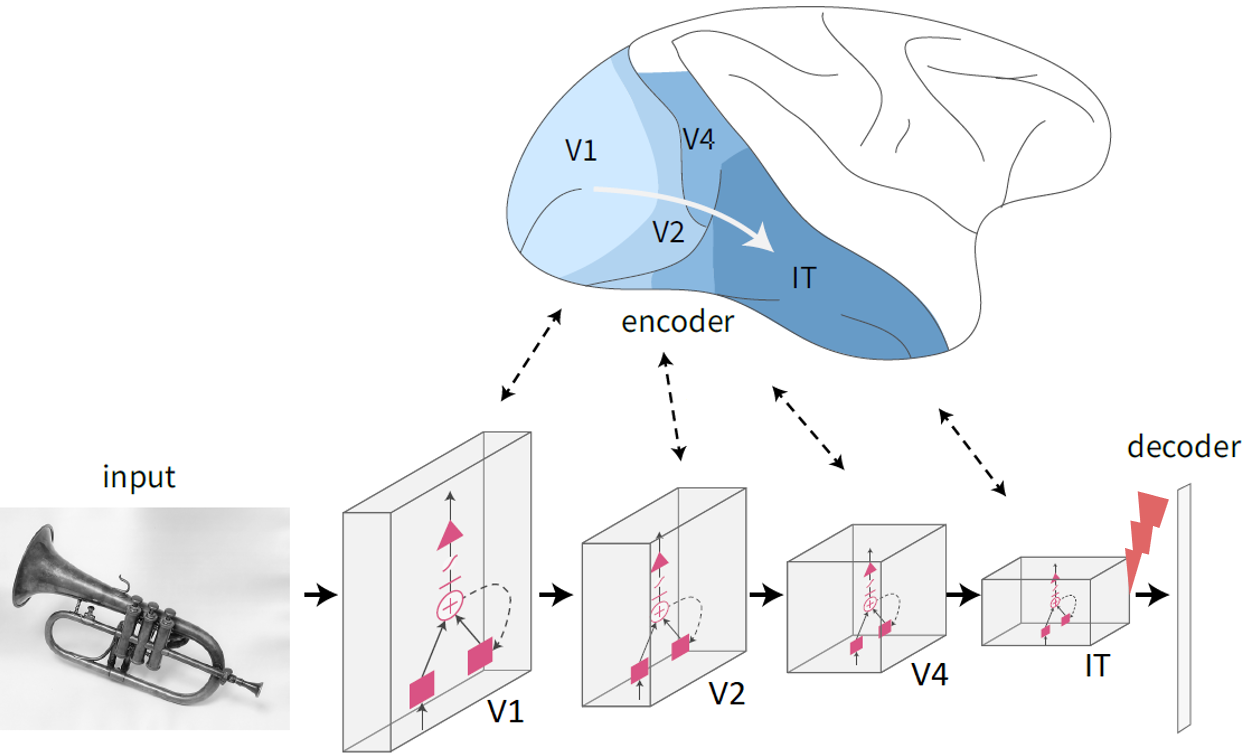 On top of that, I used the model to predict which improvements in neurotechnology hardware would best support behavioral control. I think modeling the behavioral effects of neural perturbations is an exciting direction and I’m hoping to thereby bring the models closer to clinical applications. For more details, please see the paper.
On top of that, I used the model to predict which improvements in neurotechnology hardware would best support behavioral control. I think modeling the behavioral effects of neural perturbations is an exciting direction and I’m hoping to thereby bring the models closer to clinical applications. For more details, please see the paper.
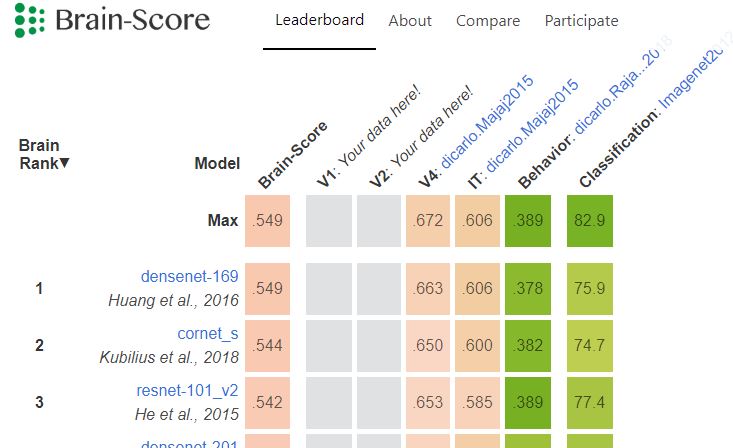
Brain-Score: Which Artificial Neural Network for Object Recognition is most Brain-Like?
The field of Machine Learning is doing pretty well at quantifying its goals and progress, yet Neuroscience is lagging behind in that regard — current claims are often qualitative and not rigorously compared with other models across a wider spectrum of tasks.
Brain-Score is our attempt to speed up progress in Neuroscience by providing a platform where models and data can compete against each other: https://www.biorxiv.org/content/early/2018/09/05/407007
Deep neural networks trained on ImageNet classification do the best on our current set of benchmarks and there is a lot of criticism about the mis-alignment between these networks and the primate ventral stream: mapping between the many layers and brain regions is unclear, the models are too large and are just static feed-forward processors.
We thus created a more brain-like model, “CORnet”, which does well on Brain-Score with only four areas and recurrent processing: https://www.biorxiv.org/content/early/2018/09/04/408385
EDIT: Science Magazine wrote a news piece about the use of deep neural networks as models of the brain with the final paragraphs devoted to Brain-Score: http://sciencemag.org/news/2018/09/smarter-ais-could-help-us-understand-how-our-brains-interpret-world
Recurrent computations for the recognition of occluded objects (humans + deep nets)
Finally out (in PNAS)! Our paper on recurrent computations for the recognition of occluded objects, in humans as well as models. Feed-forward alone doesn’t seem to cut it, but attractor dynamics help; similarly the brain requires recurrent processing to untangle highly occluded images.
http://www.pnas.org/content/early/2018/08/07/1719397115/
We have some pretty visualization gifs in the github, along with the code: https://github.com/kreimanlab/occlusion-classification
EDIT: MIT News covered our work, along with a video of us giving the intuition behind it: http://news.mit.edu/2018/mit-martin-schrimpf-advancing-machine-ability-recognize-partially-seen-objects-0920